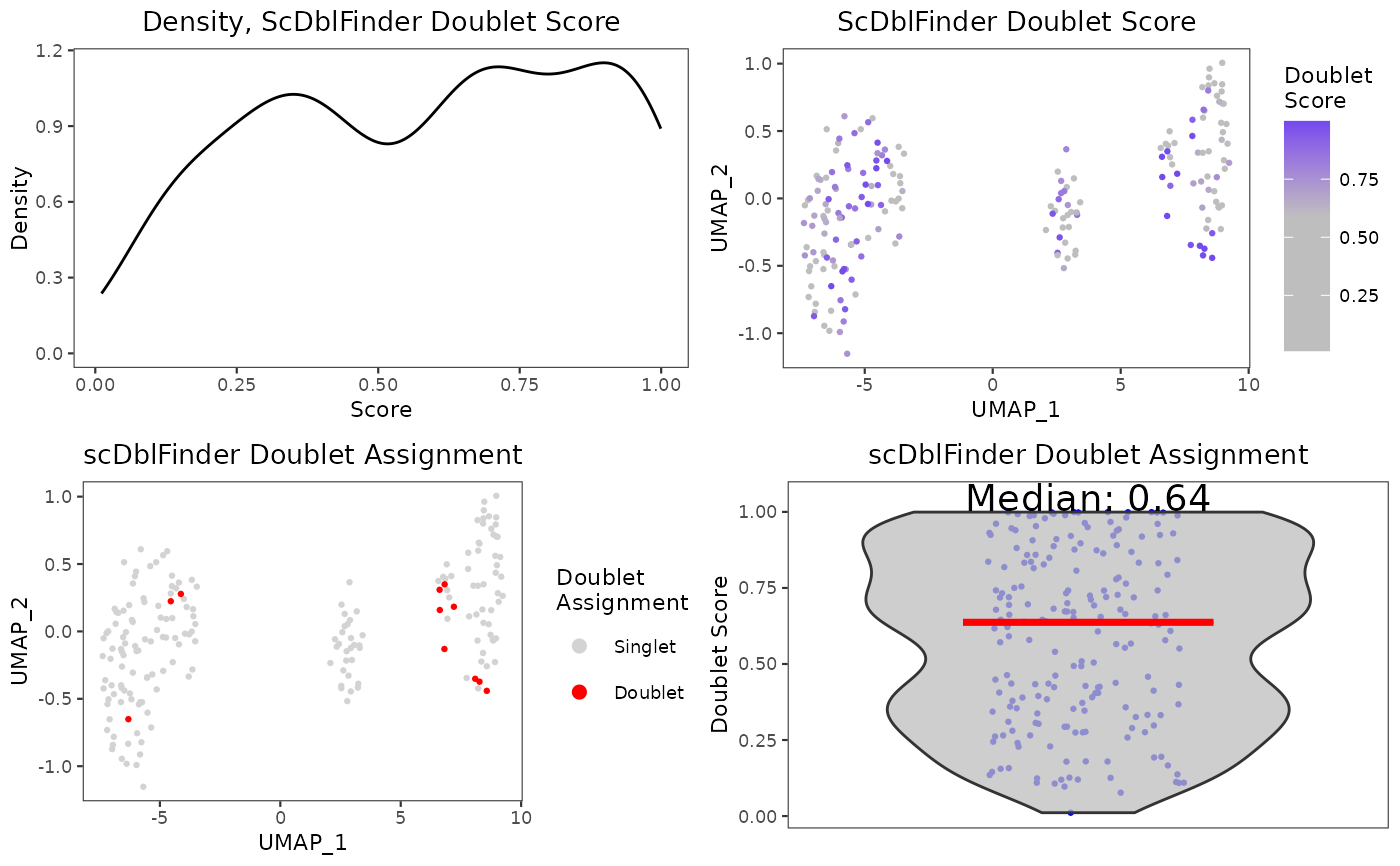
A wrapper function which visualizes outputs from the
runScDblFinder function stored in the colData slot of the
SingleCellExperiment object via various plots.
plotScDblFinderResults(
inSCE,
sample = NULL,
shape = NULL,
groupBy = NULL,
combinePlot = "all",
violin = TRUE,
boxplot = FALSE,
dots = TRUE,
reducedDimName = "UMAP",
xlab = NULL,
ylab = NULL,
dim1 = NULL,
dim2 = NULL,
bin = NULL,
binLabel = NULL,
defaultTheme = TRUE,
dotSize = 0.5,
summary = "median",
summaryTextSize = 3,
transparency = 1,
baseSize = 15,
titleSize = NULL,
axisLabelSize = NULL,
axisSize = NULL,
legendSize = NULL,
legendTitleSize = NULL,
relHeights = 1,
relWidths = c(1, 1, 1),
plotNCols = NULL,
plotNRows = NULL,
labelSamples = TRUE,
samplePerColumn = TRUE,
sampleRelHeights = 1,
sampleRelWidths = 1
)Arguments
- inSCE
Input SingleCellExperiment object with saved dimension reduction components or a variable with saved results from
runScDblFinder. Required.- sample
Character vector or colData variable name. Indicates which sample each cell belongs to. Default
NULL.- shape
If provided, add shapes based on the value. Default
NULL.- groupBy
Groupings for each numeric value. A user may input a vector equal length to the number of the samples in
inSCE, or can be retrieved from the colData slot. DefaultNULL.- combinePlot
Must be either
"all","sample", or"none"."all"will combine all plots into a single .ggplot object, while"sample"will output a list of plots separated by sample. Default"all".- violin
Boolean. If
TRUE, will plot the violin plot. DefaultTRUE.- boxplot
Boolean. If
TRUE, will plot boxplots for each violin plot. DefaultTRUE.- dots
Boolean. If
TRUE, will plot dots for each violin plot. DefaultTRUE.- reducedDimName
Saved dimension reduction name in
inSCE. Default"UMAP".- xlab
Character vector. Label for x-axis. Default
NULL.- ylab
Character vector. Label for y-axis. Default
NULL.- dim1
1st dimension to be used for plotting. Can either be a string which specifies the name of the dimension to be plotted from reducedDims, or a numeric value which specifies the index of the dimension to be plotted. Default is
NULL.- dim2
2nd dimension to be used for plotting. Similar to
dim1. Default isNULL.- bin
Numeric vector. If single value, will divide the numeric values into
bingroups. If more than one value, will bin numeric values using values as a cut point. DefaultNULL.- binLabel
Character vector. Labels for the bins created by
bin. DefaultNULL.- defaultTheme
Removes grid in plot and sets axis title size to
10whenTRUE. DefaultTRUE.- dotSize
Size of dots. Default
0.5.- summary
Adds a summary statistic, as well as a crossbar to the violin plot. Options are
"mean"or"median". DefaultNULL.- summaryTextSize
The text size of the summary statistic displayed above the violin plot. Default
3.- transparency
Transparency of the dots, values will be 0-1. Default
1.- baseSize
The base font size for all text. Default
12. Can be overwritten bytitleSize,axisSize, andaxisLabelSize,legendSize,legendTitleSize.- titleSize
Size of title of plot. Default
NULL.- axisLabelSize
Size of x/y-axis labels. Default
NULL.- axisSize
Size of x/y-axis ticks. Default
NULL.- legendSize
size of legend. Default
NULL.- legendTitleSize
size of legend title. Default
NULL.- relHeights
Relative heights of plots when combine is set. Default
1.- relWidths
Relative widths of plots when combine is set. Default
c(1, 1, 1).- plotNCols
Number of columns when plots are combined in a grid. Default
NULL.- plotNRows
Number of rows when plots are combined in a grid. Default
NULL.- labelSamples
Will label sample name in title of plot if TRUE. Default
TRUE.- samplePerColumn
If
TRUE, when there are multiple samples and combining by"all", the output .ggplot will have plots from each sample on a single column. DefaultTRUE.- sampleRelHeights
If there are multiple samples and combining by
"all", the relative heights for each plot. Default1.- sampleRelWidths
If there are multiple samples and combining by
"all", the relative widths for each plot. Default1.
Value
list of .ggplot objects
See also
Examples
data(scExample, package="singleCellTK")
sce <- subsetSCECols(sce, colData = "type != 'EmptyDroplet'")
sce <- runQuickUMAP(sce)
#> Sat Mar 18 10:29:38 2023 ... Computing Scater UMAP for sample 'pbmc_4k'.
#> Found more than one class "dist" in cache; using the first, from namespace 'BiocGenerics'
#> Also defined by ‘spam’
#> Found more than one class "dist" in cache; using the first, from namespace 'BiocGenerics'
#> Also defined by ‘spam’
#> Found more than one class "dist" in cache; using the first, from namespace 'BiocGenerics'
#> Also defined by ‘spam’
sce <- runScDblFinder(sce)
#> Sat Mar 18 10:29:41 2023 ... Running 'scDblFinder'
plotScDblFinderResults(inSCE = sce, reducedDimName = "UMAP")