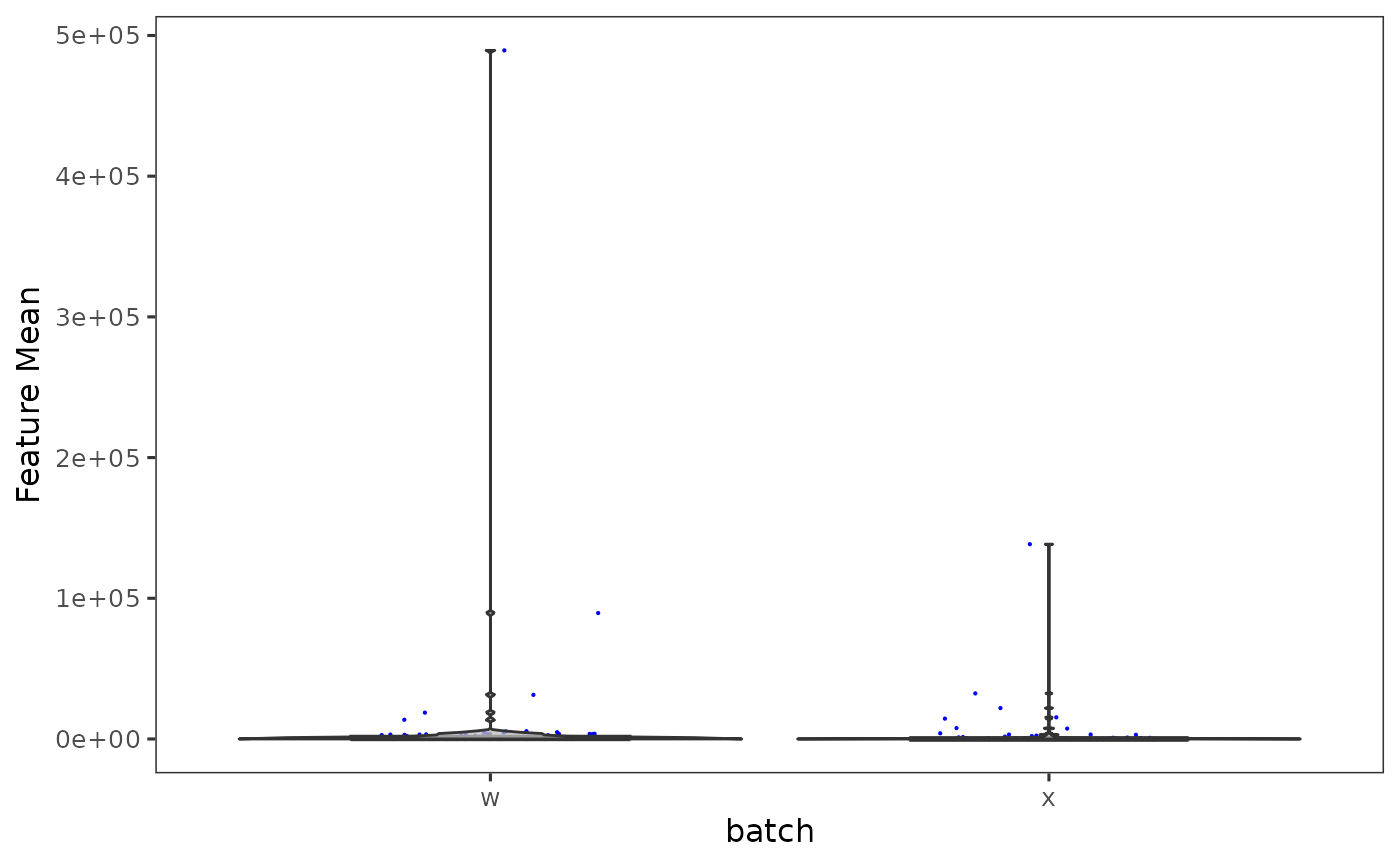
Plot mean feature value in each batch of a SingleCellExperiment object
Source:R/plotBatchVariance.R
plotSCEBatchFeatureMean.RdPlot mean feature value in each batch of a SingleCellExperiment object
plotSCEBatchFeatureMean(
inSCE,
useAssay = NULL,
useReddim = NULL,
useAltExp = NULL,
batch = "batch",
xlab = "batch",
ylab = "Feature Mean",
...
)Arguments
- inSCE
SingleCellExperiment inherited object.
- useAssay
A single character. The name of the assay that stores the value to plot. For
useReddimanduseAltExpalso. DefaultNULL.- useReddim
A single character. The name of the dimension reduced matrix that stores the value to plot. Default
NULL.- useAltExp
A single character. The name of the alternative experiment that stores an assay of the value to plot. Default
NULL.- batch
A single character. The name of batch annotation column in
colData(inSCE). Default"batch".- xlab
label for x-axis. Default
"batch".- ylab
label for y-axis. Default
"Feature Mean".- ...
Additional arguments passed to
.ggViolin.
Value
ggplot
Examples
data('sceBatches', package = 'singleCellTK')
plotSCEBatchFeatureMean(sceBatches, useAssay = "counts")