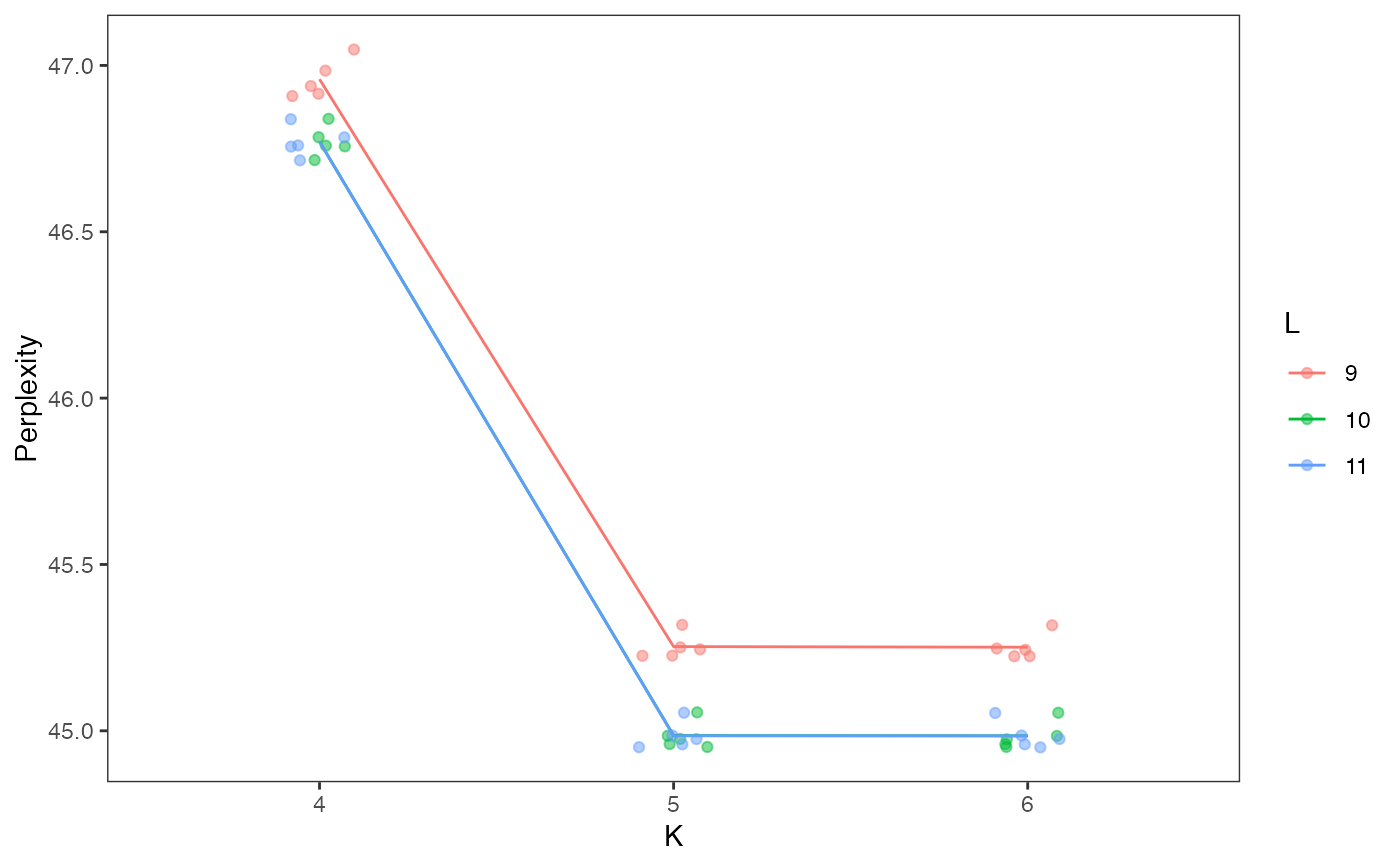
Calculate and visualize perplexity of all models in a celdaList, with count resampling
Source:R/perplexity.R
resamplePerplexity.RdCalculates the perplexity of each model's cluster assignments given the provided countMatrix, as well as resamplings of that count matrix, providing a distribution of perplexities and a better sense of the quality of a given K/L choice.
resamplePerplexity( x, celdaList, useAssay = "counts", altExpName = "featureSubset", resample = 5, seed = 12345 ) # S4 method for SingleCellExperiment resamplePerplexity( x, useAssay = "counts", altExpName = "featureSubset", resample = 5, seed = 12345 ) # S4 method for ANY resamplePerplexity(x, celdaList, resample = 5, seed = 12345)
Arguments
| x | A numeric matrix of counts or a
SingleCellExperiment returned from celdaGridSearch
with the matrix located in the assay slot under |
|---|---|
| celdaList | Object of class 'celdaList'. Used only if |
| useAssay | A string specifying which assay
slot to use if |
| altExpName | The name for the altExp slot to use. Default "featureSubset". |
| resample | Integer. The number of times to resample the counts matrix for evaluating perplexity. Default 5. |
| seed | Integer. Passed to with_seed. For reproducibility, a default value of 12345 is used. If NULL, no calls to with_seed are made. |
Value
A SingleCellExperiment object or
celdaList object with a perplexity
property, detailing the perplexity of all K/L combinations that appeared in
the celdaList's models.
Examples
data(sceCeldaCGGridSearch) sce <- resamplePerplexity(sceCeldaCGGridSearch) plotGridSearchPerplexity(sce)data(celdaCGSim, celdaCGGridSearchRes) celdaCGGridSearchRes <- resamplePerplexity( celdaCGSim$counts, celdaCGGridSearchRes ) plotGridSearchPerplexity(celdaCGGridSearchRes)